The initial deadline set for the Luzon-wide community quarantine has ended, and still the COVID-19 outbreak shows no signs of winding down. While progress has been slow and sporadic, some have found hope in the last few days; the statistics for recovery have grown, overtaking the death toll. But the true scope of the outbreak remains difficult to pin down.
In response, many have grown to see testing as a panacea: roll it out massively and everything else should follow. There’s no denying that testing plays a central role in controlling the pandemic, but there is cause to stay level-headed about what it can and can’t do.
An understanding of how these tests work, at the most fundamental level, should help not just with tempering expectations, but with refining them.
There are many ways to test for SARS-CoV-2, but, right now, the gold-standard is through reverse transcription-polymerase chain reaction (RT-PCR, for short), which detects the virus by looking for its genetic identification. Ribonucleic acid, the virus’ genetic material, will first have to be reverse-transcribed back into its complementary deoxyribonucleic acid (cDNA). How it is done differs from kit to kit. Some processes will require an additional and separate step for it, while others have found ways to do it in the same tube that will be used in the succeeding steps.
Regardless of the specifics, the method generally relies on the work of reverse transcriptase, an enzyme named after its function. The samples will then have to sit at a temperature in which the enzyme is most productive. These details will again tend to vary. For example, one kit, provisionally approved by the United States Food and Drug Administration, declares incubation at 53oC for 10 minutes. Other kits, not indicated for COVID-19, may prescribe cooler temperatures, at around 40oC.
Even after converting it into cDNA, viral genetic material will still be nearly impossible to detect in the sample as is. Maybe it’s present, but in such tiny amounts, or maybe it gets blocked from view, so to speak, by the DNA of other cells in the sample.
PCR offers a way around this. Using short stretches of free DNA, it allows lab technicians to pinpoint which squence belongs to SARS-CoV-2. These primers, as they are called, are synthesized in the lab to be extremely specific to their targets. This is why the full genome sequence of SARS-CoV-2, published online just days after official declaration of an outbreak, is crucial.
In detecting SARS-CoV-2, test kits don’t just use one set of primers. Some will target genes that are important for the virus’ structure, while others seek out sequences that encode the spike protein. This acts like a failsafe for the kit.
“This will ensure greater success in detecting the RNA of the virus in the sample,” according to Dr Mary Grace Dacuma, Assistant Professor of Zoology at the Institute of Biological Science, University of the Philippines Los Baños. Some kits even test for the “human RNase P gene as an internal control to ensure that sample screened has nucleic acid.”
Regardless of the add-ons, it’s standard for the COVID-19 kits to also target a region of the viral genome that is specific to SARS-CoV-2.

The true strength of RT-PCR is its ability to pick out a DNA sequence and amplify it across orders of magnitude. After the primers have searched and attached to their targets (the technical term is to anneal), they will serve as landing pads for another enzyme: DNA polymerase. Its job is to build upon the primer and extend it, creating a much longer strand of DNA that complements the target viral sequence.
All of this happens inside machines called thermal cyclers. As the name suggests, they are able to hit different temperature values, stay there, and then cycle through over and over.
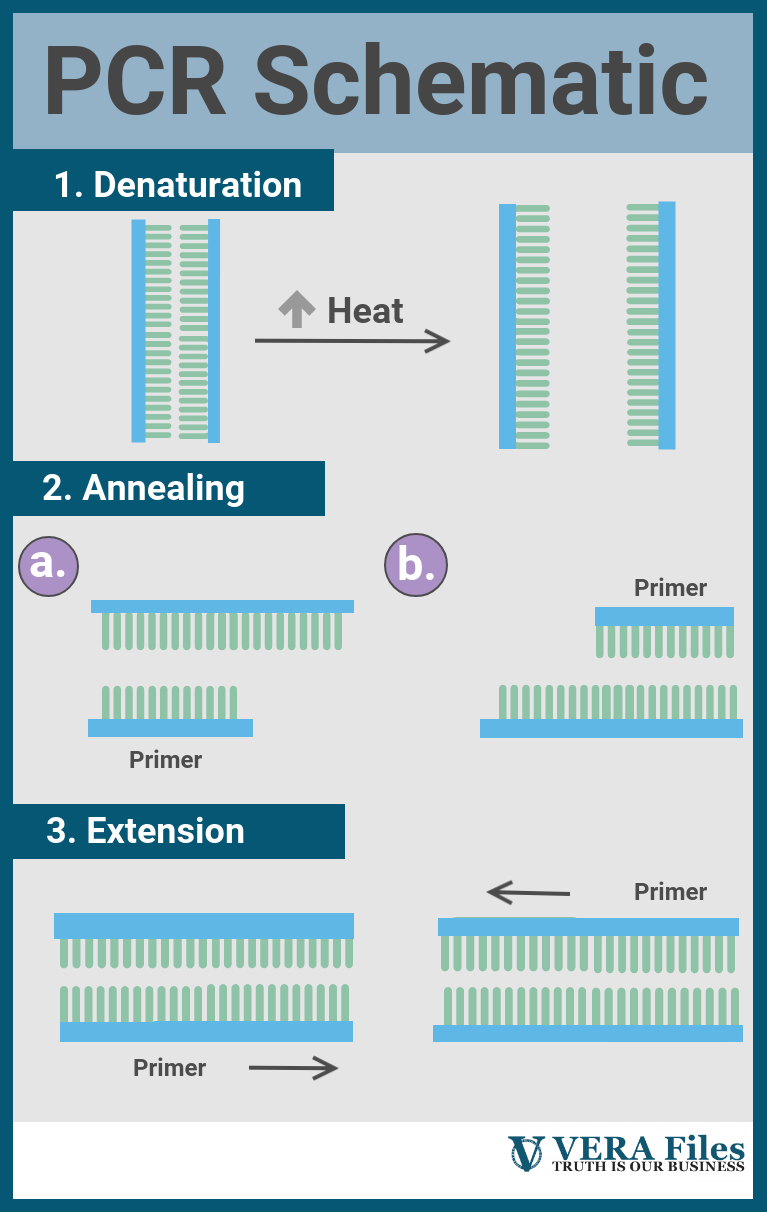
Broadly, each cycle will include three parts, each happening at a specific temperature. First, the DNA will have to be opened up to give space for the rest of the reaction. This usually requires high heat, above 90oC, but takes only a short while. After this, the thermal cycler will cool down to around 50 degrees to 60 degrees Celsius, allowing the primers to anneal with their targets.
Finally, gently turning the heat up activates the DNA polymerase and boosts its performance, letting it extend the primer efficiently.
In theory, each cycle of RT-PCR doubles the starting number of target cDNA. For example: If there is just one bit of viral cDNA in the sample to begin with, the first PCR cycle will photocopy that, so that the second cycle will have two DNA copies as its starting material. The third cycle, in turn, will start with four copies and end with eight, and so on.
Over dozens of cycles, the population of the virus cDNA in the sample, if present, grows explosively.
All of this takes place inside tubes, deep within the belly of a thermal cycler, and at a scale that is beyond the reach of the naked eye. To solve this, the tiny DNA fragments produced through RT-PCR are marked with fluorescent dyes or probes, sort of like a glow-in-the dark label.
After every RT-PCR cycle, the machine hits the tubes with light, which the fluorescent labels absorb and blast back. In the earliest stages, this may be too weak to be detected. But over time, as the cycles drag on, the machine will eventually notice the return signal, and display it as graphs in a computer monitor, tracking the exponential increase of the virus cDNA sequence in real-time.
This is what the doctors, and the medical technologists, and the lab technicians see―and what drives a COVID-19 diagnosis.
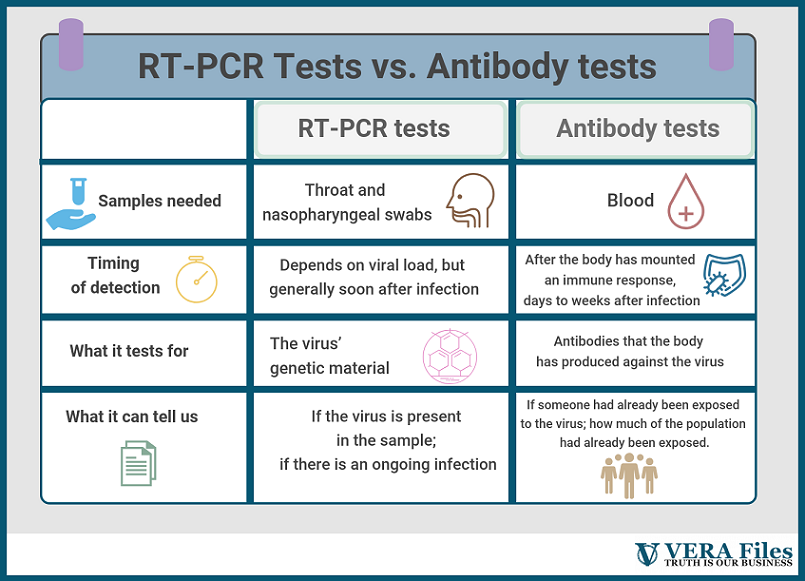
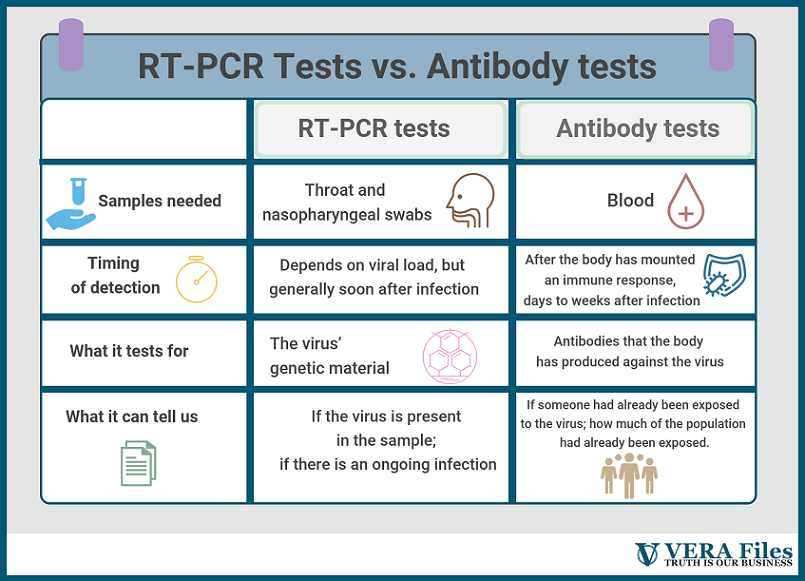
Useful as the RT-PCR tests may be, they capture an incomplete picture of the outbreak. Because they seek out the SARS-CoV-2’s genetic material, these kits can only sense an ongoing infection. After the virus has been cleared from the body, an RT-PCR test alone can’t tell whether an infection happened or not.
Serology tests, on the other hand, are made exactly for that. Instead of swabs, these use blood samples and look for antibodies, which are part of the body’s natural response to viruses. “It will not tell whether the person has an active infection or not,” Dacuma says. “It only tells that the person was exposed to the pathogen.”
This is a key distinction. RT-PCR tests can probably approximate the breadth of the problem, but using it to draw definitive conclusions about the true number of people infected is a stretch. Antibody tests have a better shot at that. In dealing with a disease that has so many concerns about asymptomatic transmission, this becomes all the more important.
One way to detect antibodies is through enzyme-linked immunosorbent assays, or ELISA, for short. This uses tiny wells coated with a protein on the virus’ surface, which fish out antibodies from the blood samples. The entire system is then visualized using a second round of antibodies that latch on to the first and carry a fluorescent or color-based label.
Unlike RT-PCR testing, this method requires that a lot more be known about the virus than just its genome: which of its structures elicits an immune response, for example, and what specific part of that do antibodies recognize. The lab work is more complicated, too, as scientists will need to artificially synthesize this structure in cells that aren’t SARS-CoV-2. The products can sometimes come out slightly deformed, unable to bind the antibodies as effectively.
Another key difference between the two tests is timing. “During the early stages of incubation period, the amount of virion in secretions or virus load in infected cells may be below the detection limit,” Dr Dacuma explains. “However, if the RT-PCR is very sensitive it may be able to detect even one copy of viral RNA and amplify this to a number detectable in the RT-PCR assay.”
Antibody tests, on the other hand, need to wait until the infection has progressed and antibodies against the virus have developed. This can take days or weeks.
Because the two tests are so different, it may be shortsighted to say that one is better than the other. Each fulfills a different role and both will be crucial in taming the contagion.
NOTE: This is a corrected version using the correct initials for complementary deoxyribonucleic acid (cDNA)





